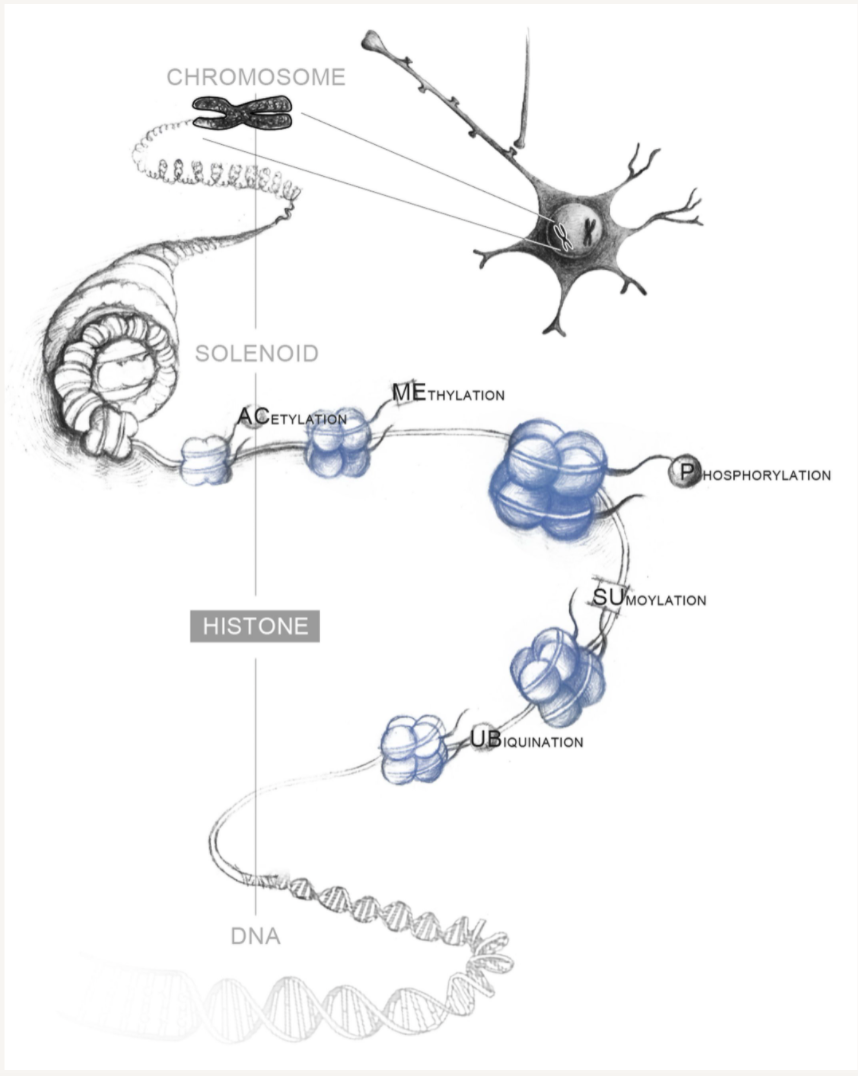
Neuronal gene regulation and chromatin.
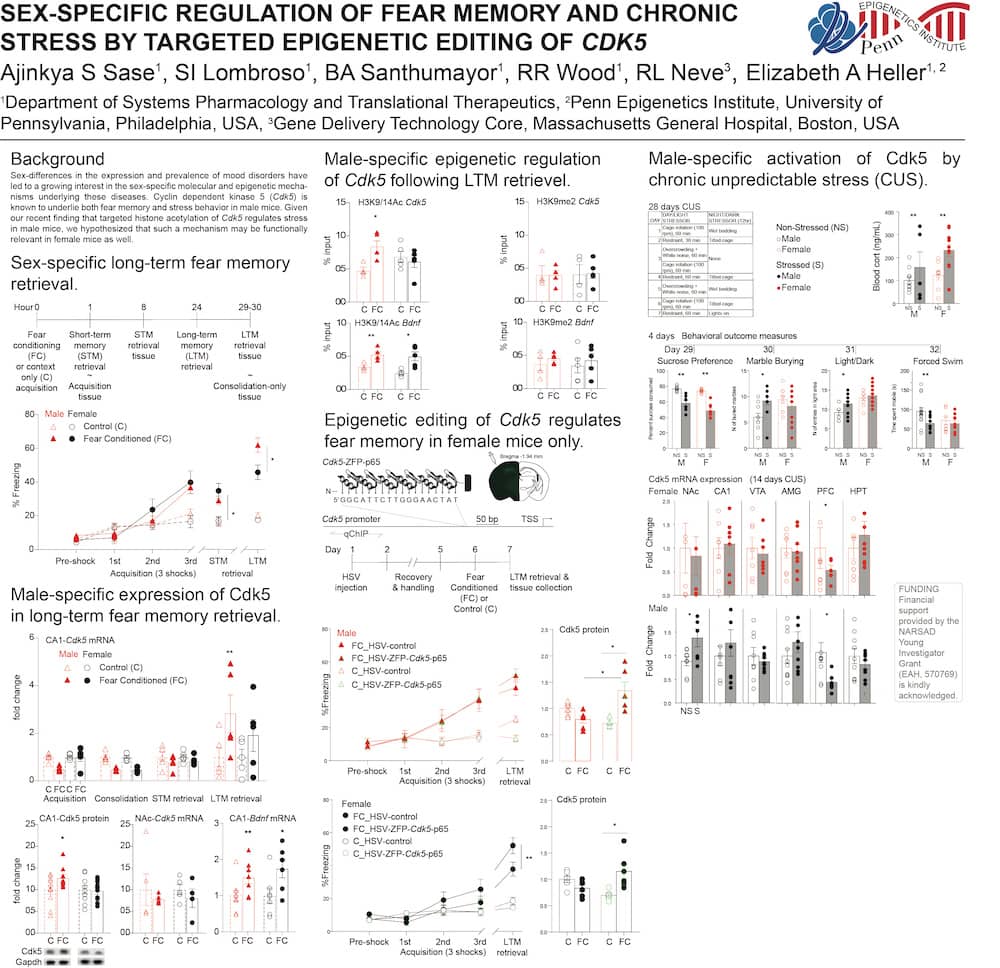
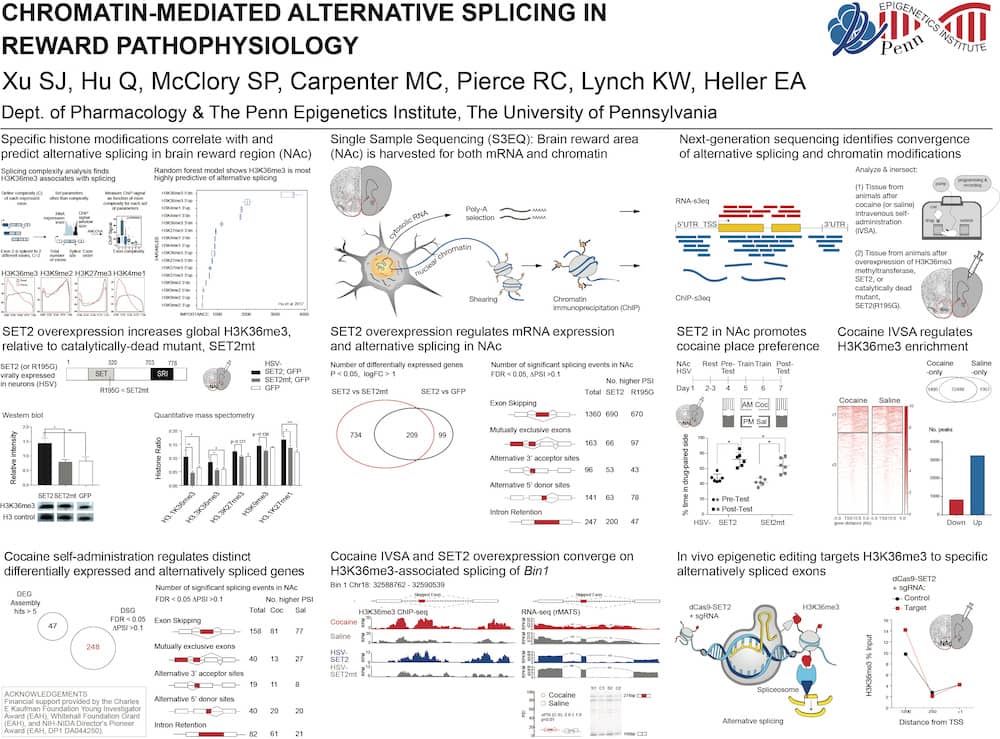
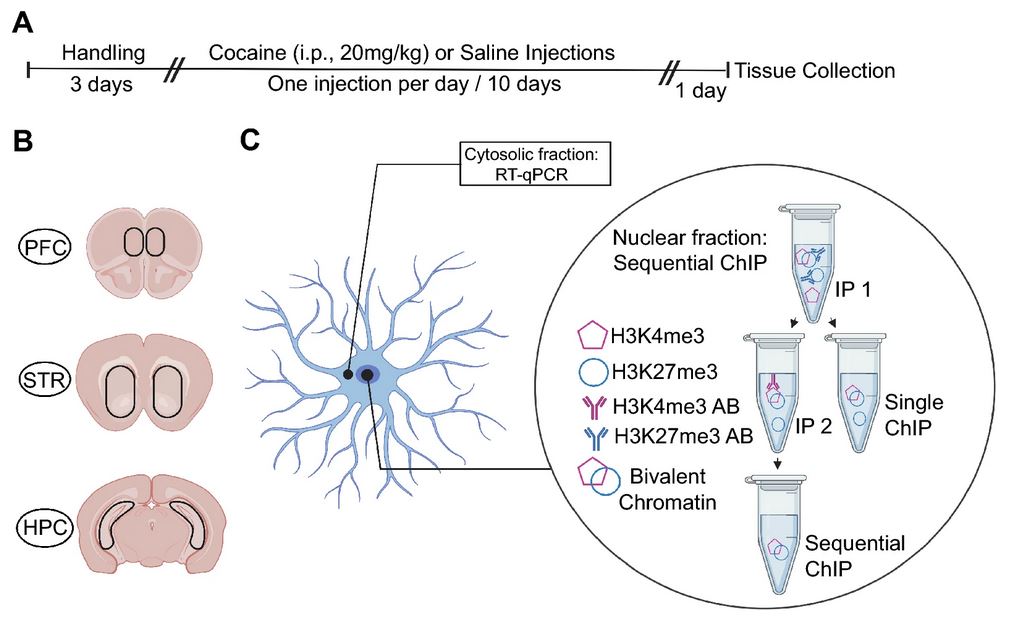
The Heller Lab studies the gene regulatory mechanisms that underlie psychiatric disease. To approach this problem, we apply preclinical mouse paradigms of drug addiction and depression to define functionally relevant histone modifications and transcription factors. Because the syndromes of addiction and depression persist long after cessation of the insult, chromatin remodeling is an attractive mechanism for such long-lasting effects and presents an intriguing target for therapeutic intervention. We use high-throughput sequencing and bioinformatic approaches, including machine learning, to identify genes at which drug- or stress-regulation of the epigenome correlates with changes in gene expression. Using novel epigenetic editing tools, we then target individual modifications and examine their causal relevance to transcription and behavior. This ‘bottom-up’ approach allows direct elucidation of the causal relevance of epigenetic remodeling in the brain.
Latest News
Our very own Kechi Holmes just won a University of Pennsylvania’s Center for Undergraduate Research and Fellowships (CURF) Research Grant Award for her work Synthesis and characterization of Cytosporone B derivatives: Small molecule activation of Nuclear Receptor 4A1, a potential target for treating cocaine use disorder. Congratulations Kechi!
The Cell and Molecular Biology (CAMB) program recently hosted a friendly basketball game between graduate students and faculty members. Our very own Keegan Krick won the tip-off!

The Heller lab attends the 39th Annual Mahoney Institute for Neuroscience Symposium: The Year of Neuroimmunology which was co-organized by Liz. Graduate student Michael Murphy and post-doc Julia Winter presented their posters as part of the symposium.


Dr. Heller is a faculty leader of the Trainee Advocacy Alliance (TAA), a collaborative group of Biomedical Graduate Studies (BGS) graduate students, Biomedical Postdoctoral Program (BPP) post-docs, and Penn faculty members who are committed to creating, nurturing, and maintaining a Penn biomedical research community that embodies inclusivity, diversity, and equity.